Source code:
Background
Plasmid analysis is a fundamental component of modern molecular biology, synthetic biology, and genetic engineering workflows. Researchers routinely use plasmids as vectors for gene cloning, protein expression, CRISPR systems, and recombinant DNA applications.
Accurate visualization of plasmid structure and restriction enzyme cut sites is essential for experimental design, construct verification, digestion planning, and sequence validation. Traditional plasmid mapping tools are often limited by proprietary licensing, lack of customization, or inflexible workflows.
As sequencing throughput and synthetic construct complexity continue to increase, there is growing demand for lightweight, programmable, and reproducible bioinformatics tools capable of integrating sequence analysis with automated visualization. PlasMap addresses this need by providing an open and extensible Python-based framework for restriction enzyme analysis and plasmid map generation, enabling streamlined plasmid characterization and graphical annotation for research and educational applications.
Description
PlasMap is a Python-based plasmid analysis and visualization tool designed to identify restriction enzyme recognition sites and generate annotated plasmid maps from DNA sequence data.
The program accepts plasmid sequences in FASTA format and scans the sequence against a user-defined or database-derived restriction enzyme set. The software detects recognition motifs on both the forward strand and reverse-complement strand, records enzyme cut positions, and organizes these features into structured annotations. Using this information, PlasMap generates both circular and linear plasmid visualizations with labeled restriction sites and positional markers.
The program is intended for molecular cloning workflows, plasmid verification, restriction digest planning, and educational demonstrations of recombinant DNA analysis. Its modular architecture also allows future expansion into additional bioinformatics capabilities such as ORF prediction, GenBank feature integration, primer design, and CRISPR annotation.
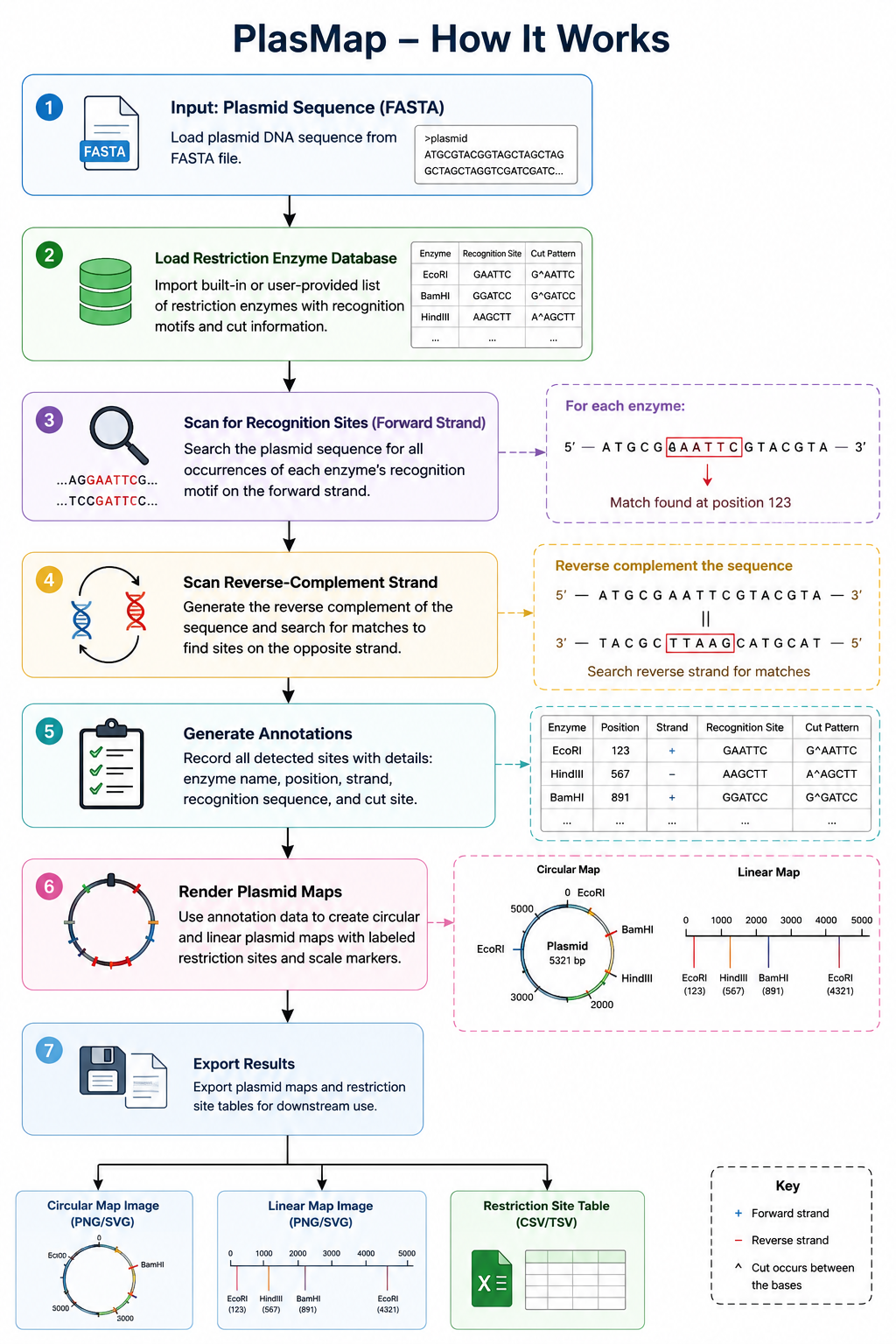
How It Works
PlasMap operates by first loading a plasmid DNA sequence from a FASTA file and validating the sequence to ensure it contains properly formatted nucleotide data.
The program then imports a restriction enzyme database containing enzyme names, recognition motifs, and cut-site information. For each enzyme, PlasMap scans the plasmid sequence to identify recognition sites on both the forward strand and the reverse-complement strand, allowing accurate detection of restriction sites regardless of orientation.
All identified matches are recorded as structured annotations that include the enzyme name, recognition sequence, strand orientation, and positional coordinates within the plasmid.
These annotations are subsequently used to generate both circular and linear plasmid maps with labeled restriction sites and scale markers:
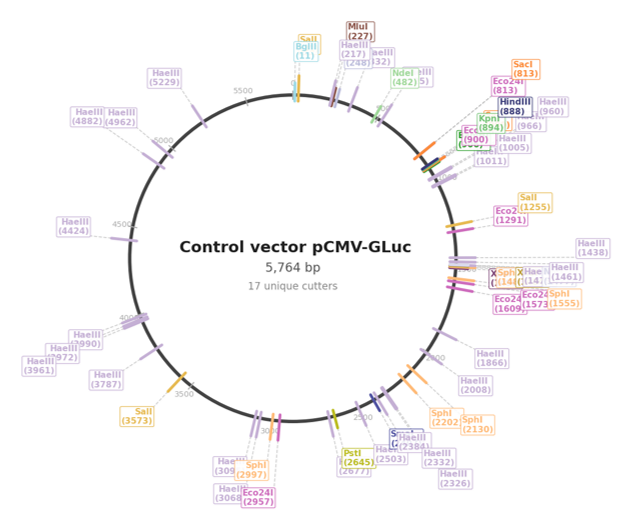
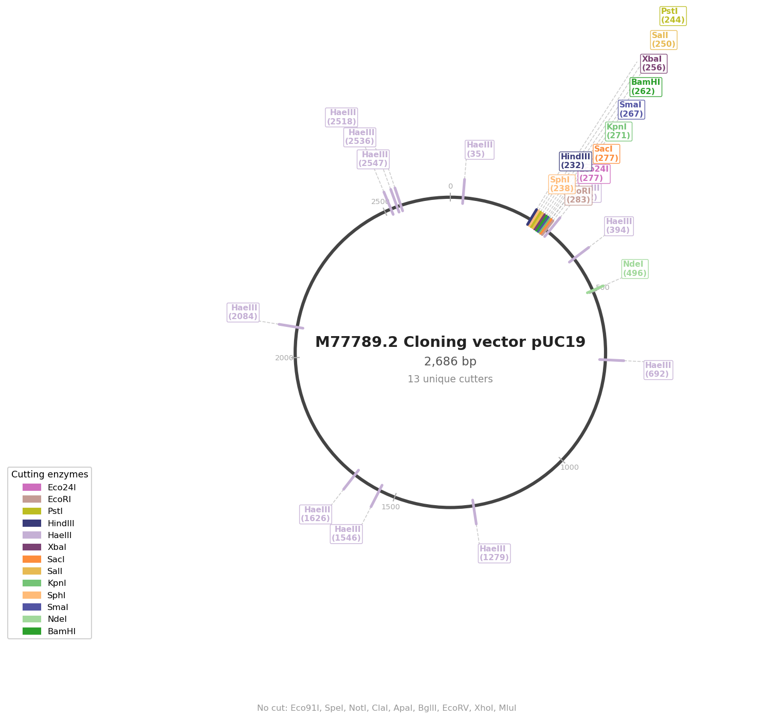
Circular Map:

Linear Map:
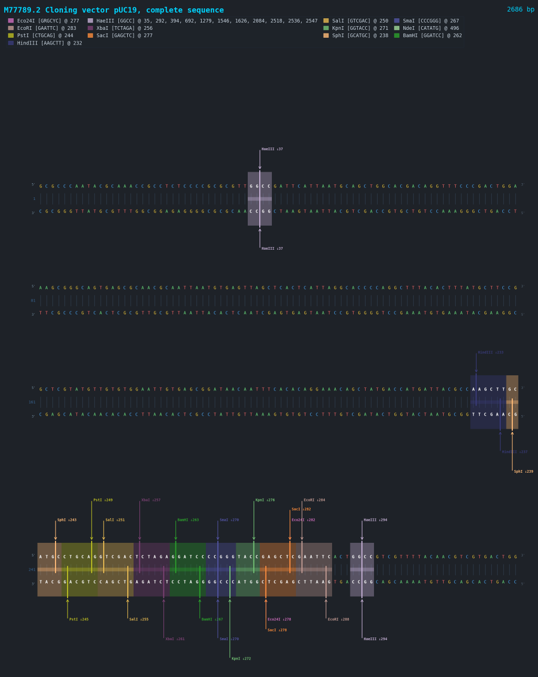
Finally, the program exports the resulting visualizations and restriction site tables in standard graphical and tabular formats, enabling downstream analysis, cloning workflow design, and publication-quality plasmid documentation.