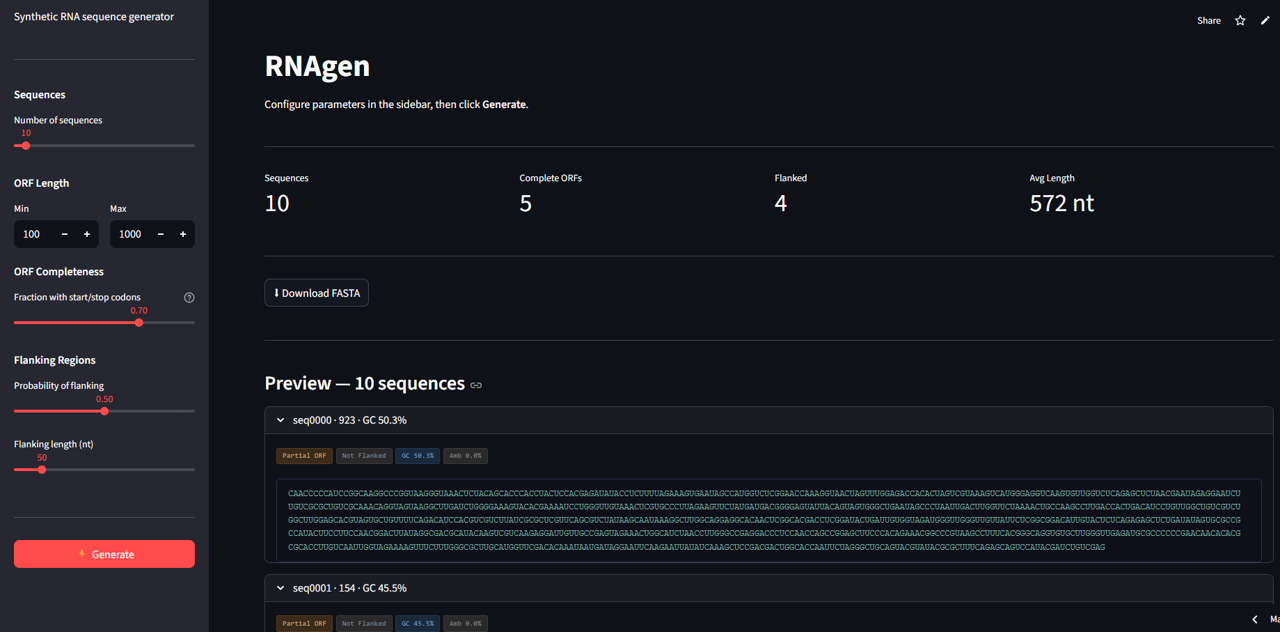
Click here for a Demo
Background:
Experimental RNA sequencing pipelines and bioinformatics tools require large volumes of test data to validate correctness, benchmark performance, and stress test edge cases — data that is often unavailable early in development or impractical to source from wet lab experiments. Sourcing real sequencing data is constrained by the cost and time of wet lab experiments, access to sequencing infrastructure, and the difficulty of obtaining sequences with specific properties on demand. Synthetic data generation fills this gap by providing reproducible, configurable datasets that can be produced instantly and tailored to the requirements of a given pipeline or experiment.
Description:
RNAgen generates synthetic RNA sequences with configurable biological properties and outputs them in standard FASTA format. Each sequence is constructed with an open reading frame that is either complete — containing an authentic AUG start codon and a UAA, UAG, or UGA stop codon — or partial, simulating incomplete or degraded transcript reads. Optional flanking regions can be added to either side of the ORF to reflect the untranslated regions present in real transcript architecture. Sequence length, ORF completeness ratio, flanking probability, and flanking length are all user-controlled, allowing targeted simulation of specific biological conditions. Output is immediately compatible with downstream tools such as BLAST, RNA-seq aligners, and ORF prediction software.
How It Works
RNAgen is built around a Simulator class that accepts user-defined parameters and orchestrates sequence generation. When a run is initiated, the class iterates over the requested number of sequences, randomly determining for each whether it will be a complete or partial ORF based on the configured completeness ratio. Complete ORFs are assembled codon by codon — starting with AUG, filling the body with randomly selected codons, and terminating with a randomly chosen stop codon. Partial ORFs bypass this structure and are generated as raw random nucleotide sequences of the specified length. Flanking regions are added probabilistically, with independent random sequences appended to the 5′ and 3′ ends of the ORF when the flanking probability threshold is met.
Each generated sequence is paired with a metadata description capturing its length, GC content, ambiguity content, ORF type, and flanking status, all computed via utility functions in sequence_lib.py. The full set of results is stored in self.results as a list of tuples containing the sequence ID, description, and sequence string. Once generation is complete, results are written to disk in FASTA format using write_fasta(). The CLI entry point in main.py handles argument parsing and validation before passing a configuration namespace to the Simulator, while app.py provides a Streamlit web interface that instantiates the same class directly, bypassing the CLI layer entirely.